SARS-CoV-2 Sequencing

SARS-CoV-2 Sequencing
- Comprehensive Next Generation Sequencing (NGS) Service
- Sequence the Entire SARS-CoV-2 Viral Genome from Various Sample Types
- Detect Novel SARS-CoV-2 Variants Utilizing COVID-19 ARTICi Amplicon Sequencing Protocol
- Illumina® MiSeq Platform
- Examine Inter-Individual and Intra-Individual Variations of the SARS-CoV-2 Virus
- Fast Turn Around Time
Norgen Biotek offers Next-Generation Sequencing (NGS) in an accredited state-of-the-art laboratory, utilizing the Illumina sequencing platforms. Our comprehensive service includes sample isolation, library preparation and sequencing to full bioinformatics analysis. We have extensive expertise in sample preparation, sequencing and analysis of all types of samples.
Background
In December of 2019, a novel strain of coronaviruses, SARS-CoV-2, was discovered leading to an outbreak of infectious disease causing a global pandemic. Coronaviruses are a large family of viruses known to infect both animals and humans. In humans, SARS-CoV-2 coronavirus infections can cause various illnesses from the common cold to more severe diseases such as Middle East Respiratory Syndrome (MERS) and Severe Acute Respiratory Syndrome (SARS).
To successfully infect a healthy cell, coronaviruses must bind to the target healthy cell by binding to specific receptors on the healthy cell membrane. The SARS-CoV-2 Spike (S) glycoprotein, an 180k-Da oligomeric S proteinii, is the single viral membrane protein which facilitates binding to the target cells which in turn meditates viral-cell fusion. While qRT-PCR based technologies can determine presence or absence of the SARS-CoV-2 coronavirus in biological samples, the importance of detecting mutations in the viral genome is becoming more and more important, as new variants of the virus emerge. The SARS-CoV-2 Spike (S) protein gene has been found to contain one of the key mutations that may influence the function of the protein in its ability to infect healthy cellsiii. In addition to the variant which arises from the mutation in Spike (S) protein gene, other mutations have now been detected throughout the viral genome. The genes containing these mutations represent the most important targets for COVID-19 vaccine and therapeutic researchiv.
Next Generation Sequencing of the SARS-CoV-2 viral genome allows for identification of critical viral variants that can provide insight to viral evolution and severity of infection. The initial ARTIC SARS-CoV-2 sequencing protocol was introduced in January 2020 and was widely adopted by many genomics institutions across many countries worldwide. In September 2020 improvements to the original protocol were published and it has since become the most popular method for amplicon sequencing of SARS-CoV-2.
i. Tyson, J. R., James, P., Stoddart, D., Sparks, N., Wickenhagen, A., Hall, G., ... & Quick, J. (2020). Improvements to the ARTIC multiplex PCR method for SARS-CoV-2 genome sequencing using nanopore. bioRxiv.
ii. Bosch, B. J., Van der Zee, R., De Haan, C. A., & Rottier, P. J. (2003). The coronavirus spike protein is a class I virus fusion protein: structural and functional characterization of the fusion core complex. Journal of virology, 77(16), 8801-8811.
iii. McNamara, R. P., Caro-Vegas, C., Landis, J. T., Moorad, R., Pluta, L. J., Eason, A. B., ... & Dittmer, D. P. (2020). High-density amplicon sequencing identifies community spread and ongoing evolution of SARS-CoV-2 in the Southern United States. Cell reports, 33(5), 108352.
iv. Huang, Y., Yang, C., Xu, Xf. et al. Structural and functional properties of SARS-CoV-2 spike protein: potential antivirus drug development for COVID-19. Acta Pharmacol Sin 41, 1141–1149 (2020). https://doi.org/10.1038/s41401-020-0485-4
Our NGS Clients
We are pleased to work with some of the most prestigious companies, in 140+ countries worldwide.
Your Custom Workflow
1. Consultation
2. Sample Submission/Shipping
3. Sample Isolation
4. DNA Quality Control
5. Library Preparation
6. Sequencing
7. Bioinformatics Analysis
8. Final Report
Step 1: Consultation
It is essential to have an initial engagement regarding your sequencing project with our experts to define and set clear objectives. We will discuss with you the various elements of your project to learn how we can best achieve your goals. Our NGS scientists and bioinformaticians will be happy to learn about your project and make the best recommendations to suit your project needs.
Please contact us to schedule your complimentary consultation.
Request Your Consultation
Step 2: Sample Submission
We will provide detailed instructions on how to handle, process and ship your samples* to our facility. We accept both swabs/saliva or purified RNA from SARS-CoV-2 Positive** samples. In addition, we offer a complete guideline on sample collection and preservation.
*minimum number of 10 samples required for sample submission
**If your samples require SARS-CoV-2 screening by qRT-PCR to determine positive/negative status, Norgen offers this service at an additional cost.
How will you submit your samples?

Step 3: Shipping
For detailed shipping instructions, please refer to the document below.
Shipping Instructions
Step 4: Sample Isolation
If you are submitting known SARS-CoV-2 positive samples collected on Viral Transport Media (VTM) or Universal Transport Media (UTM), or other preservative media, Norgen can perform the viral RNA extraction at our facility. For undetermined samples, we will first extract total RNA from the sample and then detect SARS-CoV-2 by qRT-PCR.We have tremendous experience and expertise in isolating viral RNA from ultra-low concentration samples such as saliva. Our dedicated lab technicians utilize our proprietary silicon carbide (SiC) resin for all isolations which allows us successfully to capture viral RNA with superior sensitivity.
The sensitivity of most commercially available diagnostic viral qRT-PCR detection kits are highly affected by the RNA quality that is used for viral detection. Norgen’s SiC technology yields viral RNA with the highest quality enabling the detection of as low as 400 copies/mL of saliva (ie. 10 viral copies/PCR reaction). This is ideal for the detection of ultra low viral loads particularly in asymptomatic and recovering patient samples.
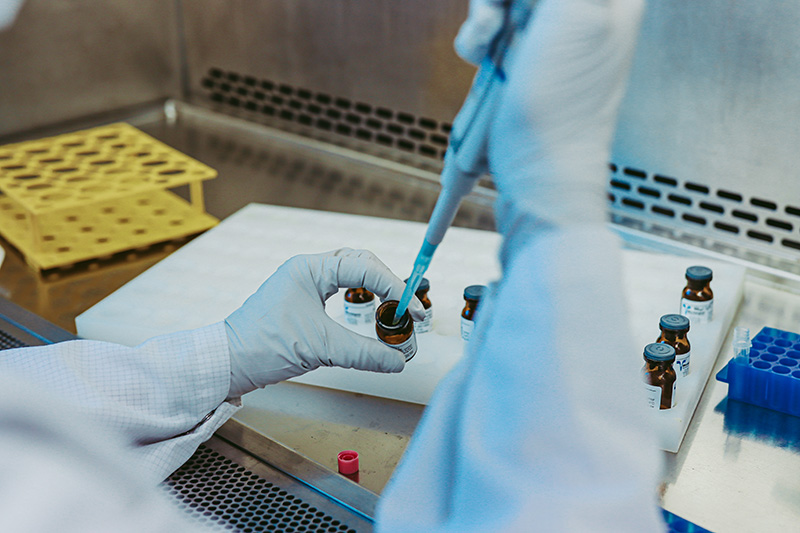

Step 5: RNA Quality Control
For undetermined samples, Norgen will first confirm SARS-CoV-2 Positive status by qRT-PCR for an additional cost.

Step 6: Library Preparation
Norgen will follow the standard library preparation procedures for SARS-CoV-2 sequencing based on the ARTIC sequencing protocol. The DNA library will then undergo a QC step prior to sequencing.
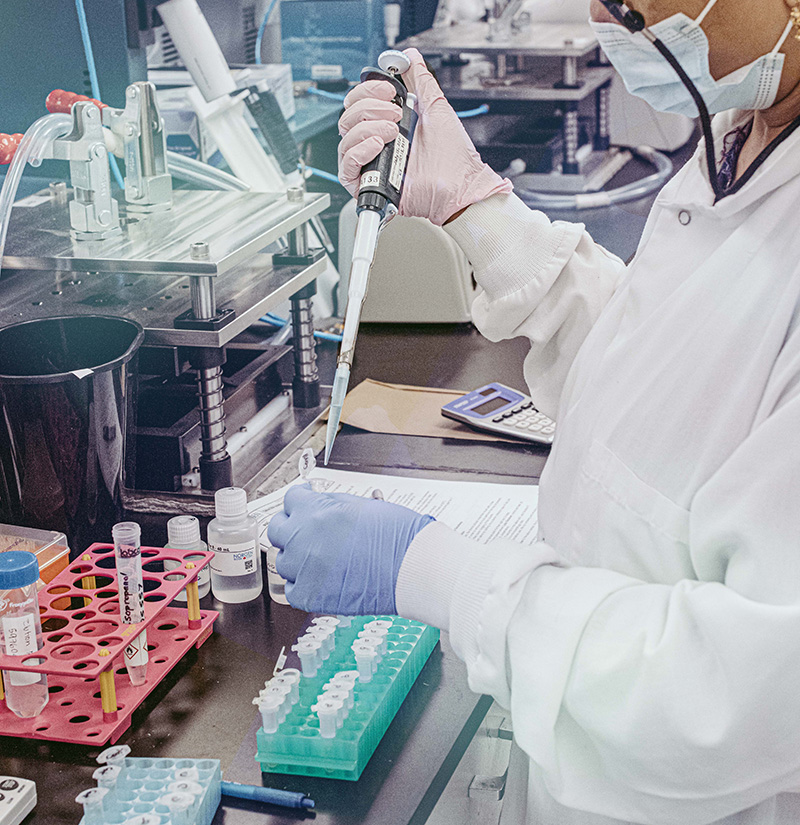

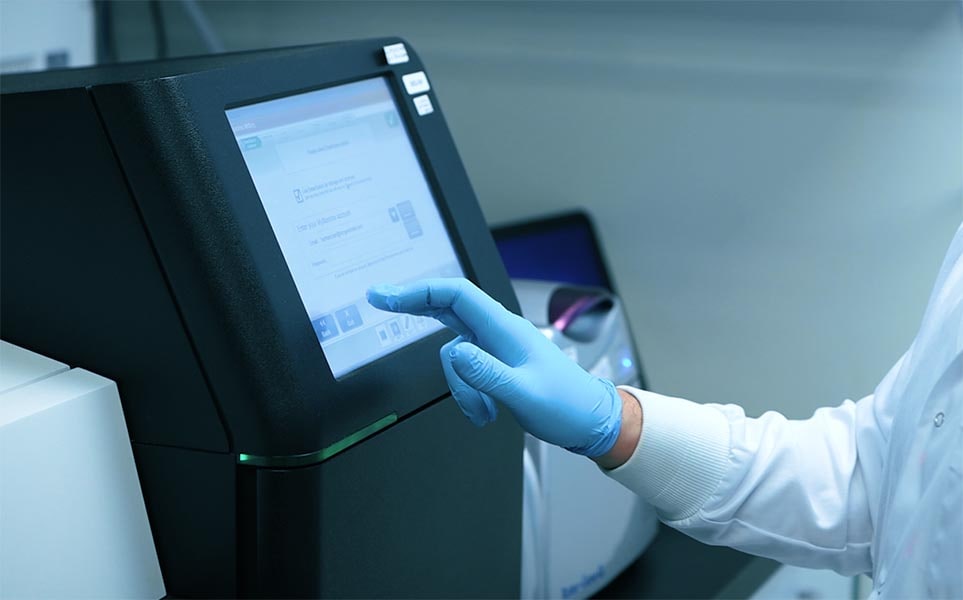
Step 7: Sequencing
We will sequence the sample libraries on the Illumina MiSeq platform to achieve your required coverage and read depth.

Step 8: Bioinformatics Analysis
Norgen has full bioinformatics capabilities from basic to advanced analysis pipelines. Our SARS-CoV-2 Sequencing service includes a full analysis to detect and identify the SARS-CoV-2 variants.
Sample Report
Let's get your project started.
Request Your ConsultationCitations
| Title | Alteration of miRNAs in Small Neuron‑Derived Extracellular Vesicles of Alzheimer's Disease Patients and the Effect of Extracellular Vesicles on Microglial Immune Responses |
| Journal | Journal of Molecular Neuroscience . 2022. |
| Authors | Devrim Yagmur Durur, Bora Tastan, Kemal Ugur Tufekci, Melis Olcum, Hamdiye Uzuner, Gökhan Karakülah, Gorsev Yener, Sermin Genc |