This is the second blog in the Movember series. Movember is the leading charity changing the face of Men's Health. Their story started 20 years ago with a few friends, some moustache's and a cheque to the Prostate Cancer Foundation of Australia. Since then they have significantly increased their fundraising efforts and impact over the years. Since 2011, Movember has committed $43 million AUD to seven key global research projects. The first project is GAP1: Prostate Cancer Biomarkers.

Biomarkers for Prostate Cancer - Beyond the PSA test
What are Biomarkers
Biomarkers can be DNA, RNA, proteins, or chemicals produced by the body that indicate the presence or severity of a disease. Biomarkers can be used as signals for early detection, to help make treatment choices, or to monitor disease progression or the response to a certain treatment. Biomarkers can be detected in any tissue or fluid and are a critical component of a personalized medicine approach to cancer treatment.
NORBLOG
Want to hear more from Norgen?
Join over 10,000 scientists, bioinformaticians, and researchers who receive our exclusive deals, industry updates, and more, directly to their inbox.
For a limited time, subscribe and SAVE 10% on your next purchase!
SIGN UP
The Need for New Biomarkers
The PSA test
The Prostate Specific Antigen (PSA) is a protein that is produced by the prostate under normal circumstances. However, in almost all cases of prostate cancer, PSA levels increase, and thus quantification of this protein is used as a biomarker for prostate cancer. Currently, the PSA test is the main method of screening for prostate cancer, which involves taking a blood sample from a patient and sending it to a laboratory where they measure the level of PSA in the blood. Although this is the best screening tool that currently exists, the sensitivity of the test is low. Age and ethnicity can affect the normal range of PSA levels, and things like urinary tract infections, excessive exercise, or recent ejaculation can also increase PSA levels. New data shows that prostate cancer treatment may be unnecessary in anywhere from 2% to 67% of cases with PSA detecting a large number of tumors that are unlikely to ever impact the patient. This is one factor pushing for the development of new biomarkers for prostate cancer.
Molecular Profiling of Tumours
Although the PSA test is the most common screening tool, a definitive diagnosis is usually made using a tissue biopsy and subsequent histological analysis. In addition to histological analysis, molecular profiling of tumors has been shown to improve the selection of personalized cancer treatment, patient responses, detection of drug resistance, and monitoring of tumor relapse.
With respect to prostate cancer, androgen deprivation therapy (ADT) is key for the medical management of patients with metastatic prostate cancer, however, a proportion of patients are resistant to some of the main ADT drugs. Specifically, ARV7 detection in cancer cells indicates that they are resistant to abiraterone and enzalutamide. Clinicians can use this biomarker to place the patient in clinical trials for new generation ADT drugs.1,2 Despite the personalized molecular information obtained from tissue biopsies, they remain highly invasive procedures and only offer a snapshot of information.
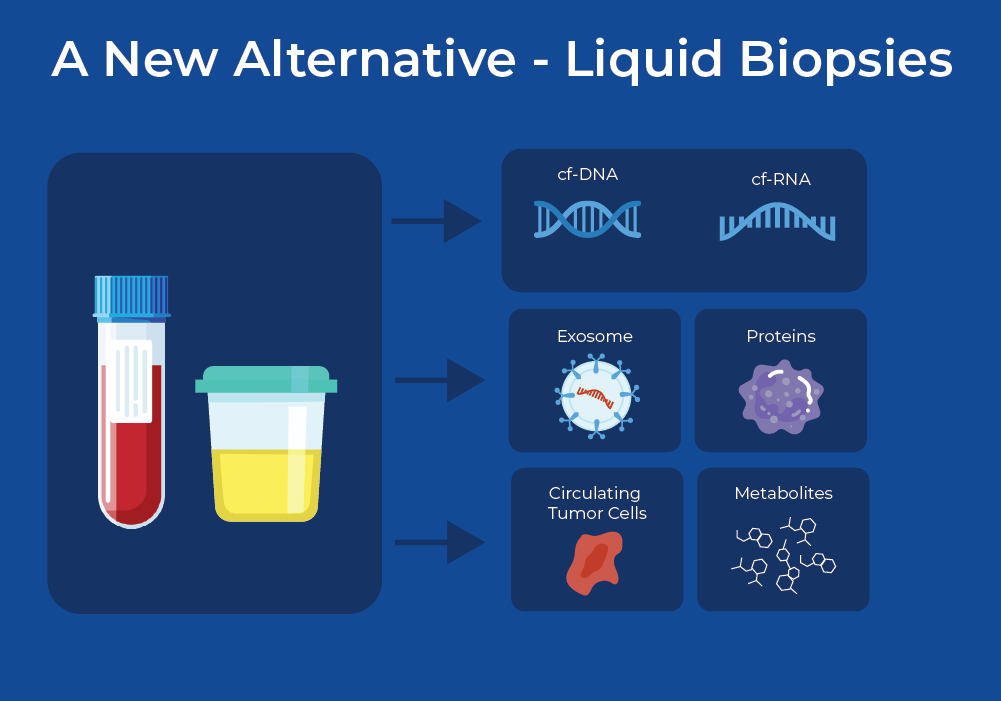
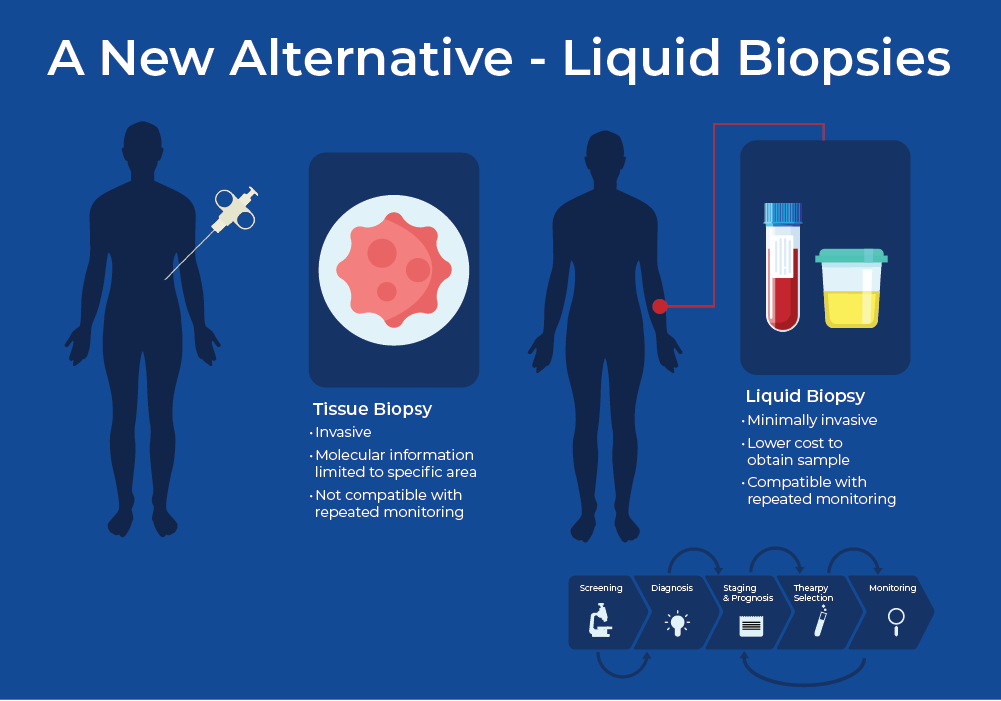
A New Alternative - Liquid Biopsies
To be able to effectively monitor response to treatments, continuous monitoring is required, and therefore the sampling procedure must be minimally invasive. Over the past decade, research into liquid biopsies has shown promise to replace tissue biopsies. Liquid biopsies offer ease of sampling and the possibility to repeatedly and less-invasively draw samples. At the moment, most liquid biopsies involve withdrawing blood to look for biomarkers, however, there is promising research concerning the use of other fluids such as urine, bile, saliva, and cerebral spinal fluid.3 These liquids harbour a variety of analytes such as DNA, RNA, and protein. These analytes can be cell-free, contained within vesicles such as an exosome, or within circulating tumour cells.
Importance of the Pre-Analytical Workflow
One of the main factors for successful cancer biomarker detection through liquid biopsies is the pre-analytical workflow. This entails analyte preservation at the site of sampling along with detailed handling protocols, transportation, and thorough sample processing.

Blood Collection and Preservation
Often, the blood cannot be immediately processed at the site where it is drawn, therefore it is critical to ensure preservation and stabilization of analytes during transportation and/or shipping. It is important to minimize leukocyte lysis as excessive genomic DNA can interfere with successful cf-DNA/RNA analysis. There are different methods for preserving the blood - Norgen tubes (Cat. 63950) rely on osmotic stabilization of nucleated cells, while other companies use a chemical crosslinking approach or biological apoptosis inhibition. Preservation devices that allow samples to be shipped at room temperature for extended periods ensure that there is minimal degradation of the biomarkers during transportation to the testing laboratory.
Recent research compared different preservation tubes and extraction methods to determine the effect on downstream analysis. This research showed that the Norgen preservation tubes yielded the most plasma and with minimal hemolysis.3 These tubes allowed for the parallel extraction of cfDNA and cfRNA from the same tube. With subsequent droplet digital PCR amplification of both cell-free DNA and RNA, they detected tumor-specific alterations in low-coverage whole-genome sequencing and DNA methylation profiling.
Urine Collection and Preservation
Blood/plasma has received most of the attention within the liquid biopsy field, however, obtaining blood samples from patients is still somewhat invasive and requires trained professionals. Using urine to detect cancer biomarkers offers a truly non-invasive sampling method that does not require trained professionals for collection. Urine samples are easily accessible, have high patient compliance, and represent the body's health due to the constant filtration of blood via the kidneys. However, issues of microbial contamination, as well as increased nucleases can compromise the analysis if the urine isn't adequately preserved.
Using Norgen's Urine Preservation tubes (Cat. 18113), urine samples were collected from 43 patients with Bladder Cancer and healthy donors. Cell-free DNA was extracted and processed with a target capture panel and deep sequencing analysis pipeline that detects genetic mutation for 118 cancers. They detected Bladder Cancer with 83.7% sensitivity and 100% specificity.4 Although this study relates to a urological cancer, novel methods designed to extract and evaluate ultra-short cfDNA may advance the liquid biopsy field for non-urological cancers as well.
In another interesting application, researchers created a modified LC/MS approach for untargeted metabolomics. They performed metabolite analysis on urine preserved in Norgen devices and detected Prostate Cancer specific metabolites.5
Nucleic Acid Extraction
Cell-free DNA/RNA
cfDNAs are degraded DNA fragments that are mainly secreted from apoptotic or malignant cells. The majority of cfDNA in healthy individuals originates from apoptotic cells with a length of 185-200 base pairs. However, in cancer patients, the levels of cfDNA are higher and fragment lengths are more uneven. A recent systematic review and meta-analysis of cell-free DNA as a diagnostic biomarker in Prostate Cancer shows favourable specificity but unsatisfactory sensitivity. However, with increased research and more specific analysis from cf-DNA, such as epigenetic or fragmentomic studies, cf-DNA still holds promise as an accurate biomarker for Prostate Cancer (PCa).6
The detection of circulating microRNA (miRNA)-based biomarkers represents another biomarker for the early detection of cancer. However, the low concentration of miRNAs released in body fluids have limited their clinical use as effective cancer biomarkers. To address this issue, a recent study showed that ultrasound treatment can amplify the release of extracellular cancer biomarkers. They showed that sonoporation amplifies the release of miRNAs from both androgen-dependent (AD) and -independent (AI) PCa cells. They identified four Prostate Cancer related miRNAs whose levels increased following ultrasound treatment and are upregulated in PCa patients compared to controls, thus confirming their clinical relevance. This highlights the potential of using ultrasound to identify novel cell-free miRNAs released from cancer cells, with the aim of developing new biomarkers with diagnostic and predictive value.7
Exosomal DNA/RNA
Exosomes are 40-150 nm-sized extracellular vesicles released by all cell types including tumor cells. Exosomes are stable in body fluids due to their lipid bilayer member and often contain DNA, RNA, and proteins. They are important in extracellular communication, and can traverse the body to localize to a specific cell. Once localized, it will fuse with the membrane of the target cell and release its contents. Often, this released cargo contributes to tumor progression. In fact, it is well-established that tumors, and even other types of diseases, release greater numbers of exosomes compared to healthy cells, which alone may be indicative of tumor burden or presence of disease.
Researchers profiled exosomal serum miRNAs (ex-miRNA) from a variety of individuals, ranging from healthy controls, those with benign prostatic hyperplasia (BPH), and others with aggressive Prostate Cancer. They performed subsequent miRNA profiling using Nanostring nCounter and identified several differentially expressed miRNAs. One of which, a tumour suppressing miR-1246, was downregulated in Prostate Cancer clinical tissues and cell lines. Thus, miR-126 offers promise as a Prostate Cancer biomarker with diagnostic potential that can predict disease aggressiveness.8
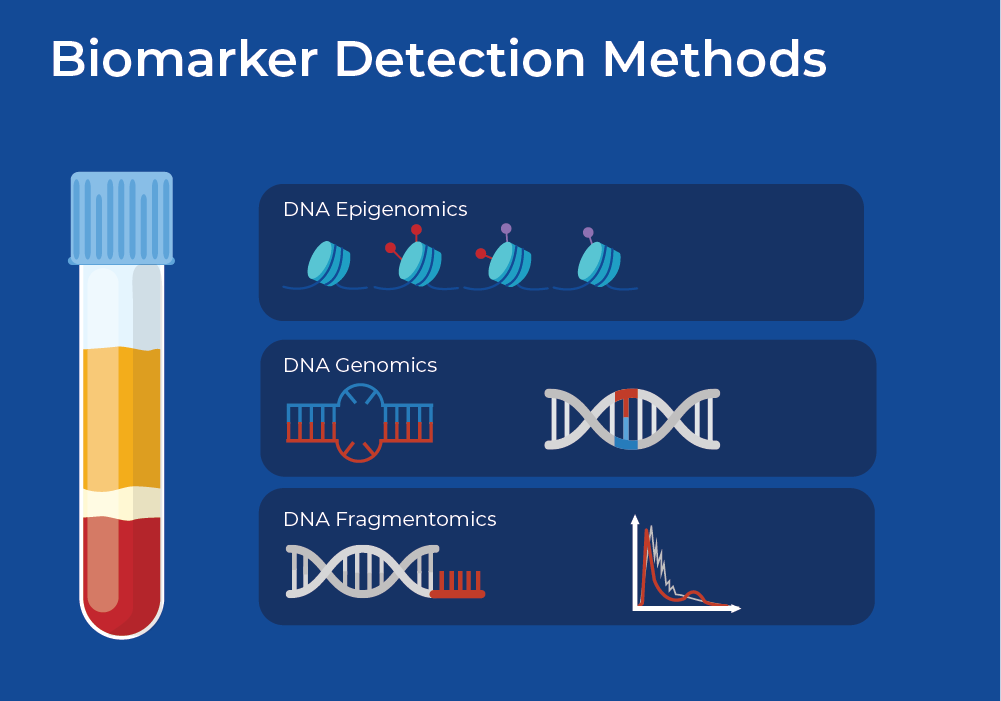
Biomarker Detection Methods
Genomics
There are specific inherited, or germline, variants that can predispose individuals to a higher risk of developing cancer. Rare variants with high penetrance are good biomarkers for cancer risk assessment. For example, it is well-known that BRCA1 and BRCA2 high-penetrance variants are strongly linked to breast and ovarian cancer. In addition, men who carry germline BRCA1/2 mutations are not only at increased risk of developing prostate cancer, but are also at risk of a more aggressive prostate cancer phenotype.9
Somatic variations are also important cancer biomarkers. Genomic instability is an important feature of cancer cells. As they adapt to the rapidly changing microenvironment, most cancers accumulate somatic mutations. These genetic alterations can range from large structural abnormalities (such as translocation, copy number alterations, or chromothripsis) to single base pair changes. These mutations can be detected in tumour tissue, as well as cell-free DNA originating from cancer cell lysis/death or active secretion, which is referred to as circulating tumor DNA (ctDNA).
Epigenomics
Epigenetic variation causes changes in DNA methylation, or histone protein modifications, without affecting the coding sequence of DNA. However, these variations affect gene and protein expression and can act as useful cancer biomarkers. DNA methlyation is an especially important modification as a loss of DNA methylation is common in many tumors and is associated with genomic instability and damage.
The detection of DNA-methylation-based biomarkers in liquid biopsies may have advantages compared to mutation detection. These include higher sensitivity and specificity in detecting early stages of cancer and in detecting residual disease, as these changes occur early on and are tissue specific. Chen et. al., recently developed a plasma DNA methylation panel "PanSeer", comprising 477 differentially methylated regions (10,613 CpG sites). They tested it with a large cohort of over 120,000 individuals, many of whom were asymptomatic for cancer. They followed them for 4 years after the blood draw and 191 individuals were later diagnosed with stomach, esophageal, colorectal, lung, or liver cancer. They showed that PanSeer detects five common types of cancer in 95% of asymptomatic individuals up to 4 years prior to diagnosis.10
Fragmentomics
Within the liquid biopsy cell-free DNA research field, a new approach called Fragmentomics is gaining traction. Fragmentomic analysis examines the fragmentation patterns of the entire cfDNA population, and is not constrained to the detection of relatively few specific mutations. This Fragmentomic pattern is non-random and represents epigenetic regulation. Many of the resultant properties of the fragment pool can be used to detect cancer, and even the origin tissue. These properties include : 1) fragment length distribution, 2) End-motif sequences, 3) Nucleosome footprint.
In a paper published last month, researchers developed novel patient-derived organoids from normal lung, normal gastric, and gastric cancer tissue. They then extracted cfDNA from the culture medium of these organoids using Norgen's Plasma/Serum Cell-Free Circulating DNA Purification Mini Kit (Cat. 55100) in both proliferative and apoptotic states, and performed fragmentomic analysis.11 Fragmentomic analysis revealed differences between cells in apoptosis versus those in proliferation, including that proliferative tissues were enriched for short fragments. This study highlights the utility of in vitro organoid models as a valuable tool for studying cfDNA biology and its associated fragmentation patterns.
Specific to prostate cancer, Helzer et. al. performed fragmentomic analysis of cfDNA from plasma. To alleviate the costs associated with whole genome sequencing, they analyzed fragmentation patterns covering coding regions from targeted panels. They used machine learning models of the fragmentation patterns to distinguish between cancer and non-cancer patients, as well as the specific tumor type and subtype. Interestingly, the classification model worked best for prostate cancer, highest ROC AUC, suggesting fragmentomic analysis of cf-DNA as a promising method for prostate cancer screening.12